Single-channel CSC With Lateral Inhibition / No Self Inhibition¶
This example demonstrates solving a convolutional sparse coding problem with a greyscale signal
where \(\mathbf{d}_{m}\) is the \(m^{\text{th}}\) dictionary
filter, \(\mathbf{x}_{m}\) is the coefficient map corresponding to
the \(m^{\text{th}}\) dictionary filter, \(\mathbf{s}\) is the
input image, and \(\boldsymbold{\omega}^T_m\) and
\(\mathbf{z}^T_m\) are inhibition weights corresponding to lateral
and self inhibition, respectively (see
admm.cbpdnin.ConvBPDNInhib).
from __future__ import print_function
from builtins import input
import numpy as np
from sporco import util
from sporco import signal
from sporco import plot
plot.config_notebook_plotting()
import sporco.metric as sm
from sporco.admm import cbpdnin
Load example image.
img = util.ExampleImages().image('kodim23.png', scaled=True, gray=True,
idxexp=np.s_[160:416, 60:316])
Highpass filter example image.
npd = 16
fltlmbd = 10
sl, sh = signal.tikhonov_filter(img, fltlmbd, npd)
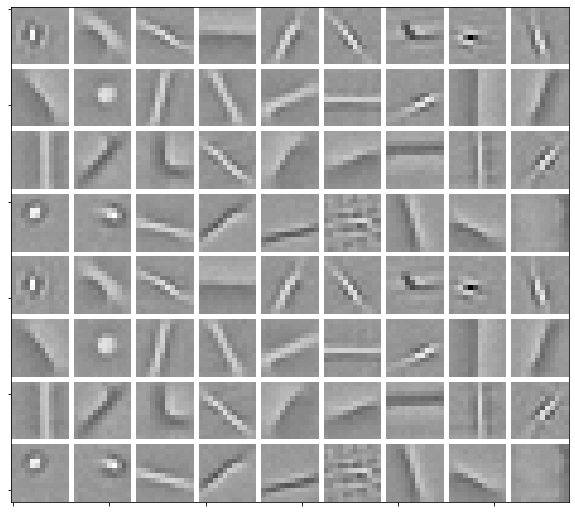
Load dictionary and display it.
D = util.convdicts()['G:12x12x36']
# Repeat the dictionary twice, adding noise to each respective pair
D = np.append(D + 0.01 * np.random.randn(*D.shape),
D + 0.01 * np.random.randn(*D.shape), axis=-1)
plot.imview(util.tiledict(D), fgsz=(10, 10))

Set admm.cbpdnin.ConvBPDNInhib solver options.
lmbda = 5e-2
mu = 5e-2
opt = cbpdnin.ConvBPDNInhib.Options({'Verbose': True, 'MaxMainIter': 200,
'RelStopTol': 5e-3, 'AuxVarObj': False})
Initialise and run CSC solver.
# Create the Ng x M grouping matrix, where Ng is the number of groups,
# and M is the number of dictionary elements. A non-zero entry at
# Wg(n, m), means that element m belongs to group n. Our dictionary
# was repeated contiguously, so elements i and i + 36 are paired for
# i = 0, ..., 35.
Wg = np.append(np.eye(36), np.eye(36), axis=-1)
# We additionally choose a rectangular inhibition window of sample
# diameter 12.
b = cbpdnin.ConvBPDNInhib(D, sh, Wg, 12, ('boxcar'),
lmbda, mu, None, opt, dimK=0)
X = b.solve()
print("ConvBPDN solve time: %.2fs" % b.timer.elapsed('solve'))
Itn Fnc DFid Regℓ1 RegLat RegSelf r s ρ
------------------------------------------------------------------------------------
0 7.62e+01 2.35e-01 1.50e+03 2.19e+01 0.00e+00 9.81e-01 6.20e-02 3.50e+00
1 6.93e+01 1.38e+00 1.32e+03 3.59e+01 0.00e+00 8.53e-01 2.06e-01 3.50e+00
2 6.01e+01 2.68e+00 1.10e+03 4.53e+01 0.00e+00 4.24e-01 2.85e-01 5.98e+00
3 5.92e+01 3.46e+00 1.05e+03 6.95e+01 0.00e+00 2.75e-01 2.35e-01 5.98e+00
4 5.83e+01 3.73e+00 1.01e+03 8.56e+01 0.00e+00 2.38e-01 1.68e-01 5.43e+00
5 5.24e+01 4.05e+00 8.81e+02 8.66e+01 0.00e+00 2.05e-01 1.38e-01 5.43e+00
6 4.89e+01 4.43e+00 8.02e+02 8.71e+01 0.00e+00 1.79e-01 1.08e-01 5.43e+00
7 4.71e+01 4.77e+00 7.56e+02 9.01e+01 0.00e+00 1.48e-01 8.85e-02 5.43e+00
8 4.54e+01 5.00e+00 7.14e+02 9.46e+01 0.00e+00 1.21e-01 7.86e-02 5.43e+00
9 4.43e+01 5.13e+00 6.84e+02 9.82e+01 0.00e+00 1.02e-01 7.12e-02 5.43e+00
10 4.35e+01 5.21e+00 6.65e+02 1.00e+02 0.00e+00 8.88e-02 6.17e-02 5.43e+00
11 4.23e+01 5.28e+00 6.39e+02 1.00e+02 0.00e+00 7.86e-02 5.34e-02 5.43e+00
12 4.07e+01 5.37e+00 6.07e+02 9.95e+01 0.00e+00 6.94e-02 4.94e-02 5.43e+00
13 3.94e+01 5.46e+00 5.81e+02 9.90e+01 0.00e+00 6.13e-02 4.60e-02 5.43e+00
14 3.89e+01 5.53e+00 5.67e+02 9.94e+01 0.00e+00 5.48e-02 4.10e-02 5.43e+00
15 3.83e+01 5.58e+00 5.54e+02 9.95e+01 0.00e+00 4.91e-02 3.74e-02 5.43e+00
16 3.75e+01 5.62e+00 5.40e+02 9.87e+01 0.00e+00 4.42e-02 3.53e-02 5.43e+00
17 3.70e+01 5.64e+00 5.29e+02 9.76e+01 0.00e+00 4.03e-02 3.30e-02 5.43e+00
18 3.64e+01 5.66e+00 5.19e+02 9.65e+01 0.00e+00 3.70e-02 3.09e-02 5.43e+00
19 3.58e+01 5.69e+00 5.07e+02 9.50e+01 0.00e+00 3.40e-02 2.99e-02 5.43e+00
20 3.54e+01 5.70e+00 5.00e+02 9.36e+01 0.00e+00 3.38e-02 2.87e-02 4.87e+00
21 3.53e+01 5.71e+00 5.00e+02 9.24e+01 0.00e+00 3.39e-02 2.70e-02 4.44e+00
22 3.52e+01 5.72e+00 5.00e+02 9.09e+01 0.00e+00 3.21e-02 2.57e-02 4.44e+00
23 3.50e+01 5.71e+00 4.96e+02 8.88e+01 0.00e+00 3.04e-02 2.49e-02 4.44e+00
24 3.46e+01 5.70e+00 4.92e+02 8.65e+01 0.00e+00 2.89e-02 2.45e-02 4.44e+00
25 3.43e+01 5.69e+00 4.88e+02 8.40e+01 0.00e+00 2.77e-02 2.41e-02 4.44e+00
26 3.40e+01 5.68e+00 4.85e+02 8.15e+01 0.00e+00 2.85e-02 2.35e-02 4.00e+00
27 3.37e+01 5.66e+00 4.82e+02 7.89e+01 0.00e+00 2.73e-02 2.31e-02 4.00e+00
28 3.35e+01 5.65e+00 4.80e+02 7.63e+01 0.00e+00 2.64e-02 2.24e-02 4.00e+00
29 3.33e+01 5.64e+00 4.79e+02 7.37e+01 0.00e+00 2.70e-02 2.15e-02 3.65e+00
30 3.31e+01 5.63e+00 4.79e+02 7.10e+01 0.00e+00 2.62e-02 2.09e-02 3.65e+00
31 3.29e+01 5.62e+00 4.78e+02 6.81e+01 0.00e+00 2.52e-02 2.04e-02 3.65e+00
32 3.27e+01 5.60e+00 4.77e+02 6.50e+01 0.00e+00 2.44e-02 1.99e-02 3.65e+00
33 3.25e+01 5.59e+00 4.76e+02 6.21e+01 0.00e+00 2.34e-02 1.94e-02 3.65e+00
34 3.23e+01 5.57e+00 4.75e+02 5.91e+01 0.00e+00 2.25e-02 1.87e-02 3.65e+00
35 3.21e+01 5.56e+00 4.74e+02 5.64e+01 0.00e+00 2.17e-02 1.81e-02 3.65e+00
36 3.18e+01 5.54e+00 4.72e+02 5.36e+01 0.00e+00 2.07e-02 1.76e-02 3.65e+00
37 3.16e+01 5.53e+00 4.70e+02 5.10e+01 0.00e+00 2.13e-02 1.70e-02 3.33e+00
38 3.14e+01 5.52e+00 4.69e+02 4.86e+01 0.00e+00 2.05e-02 1.66e-02 3.33e+00
39 3.12e+01 5.51e+00 4.68e+02 4.61e+01 0.00e+00 1.97e-02 1.60e-02 3.33e+00
40 3.10e+01 5.50e+00 4.67e+02 4.38e+01 0.00e+00 1.90e-02 1.55e-02 3.33e+00
41 3.09e+01 5.49e+00 4.66e+02 4.15e+01 0.00e+00 1.82e-02 1.48e-02 3.33e+00
42 3.07e+01 5.48e+00 4.64e+02 3.93e+01 0.00e+00 1.76e-02 1.43e-02 3.33e+00
43 3.05e+01 5.47e+00 4.63e+02 3.72e+01 0.00e+00 1.70e-02 1.39e-02 3.33e+00
44 3.03e+01 5.46e+00 4.61e+02 3.52e+01 0.00e+00 1.64e-02 1.36e-02 3.33e+00
45 3.01e+01 5.46e+00 4.59e+02 3.33e+01 0.00e+00 1.58e-02 1.31e-02 3.33e+00
46 2.99e+01 5.45e+00 4.57e+02 3.15e+01 0.00e+00 1.52e-02 1.28e-02 3.33e+00
47 2.97e+01 5.45e+00 4.56e+02 2.99e+01 0.00e+00 1.47e-02 1.24e-02 3.33e+00
48 2.96e+01 5.44e+00 4.54e+02 2.84e+01 0.00e+00 1.43e-02 1.21e-02 3.33e+00
49 2.94e+01 5.44e+00 4.53e+02 2.70e+01 0.00e+00 1.47e-02 1.19e-02 3.04e+00
50 2.93e+01 5.43e+00 4.53e+02 2.56e+01 0.00e+00 1.43e-02 1.16e-02 3.04e+00
51 2.93e+01 5.43e+00 4.52e+02 2.44e+01 0.00e+00 1.38e-02 1.11e-02 3.04e+00
52 2.92e+01 5.42e+00 4.52e+02 2.33e+01 0.00e+00 1.35e-02 1.08e-02 3.04e+00
53 2.91e+01 5.42e+00 4.52e+02 2.21e+01 0.00e+00 1.31e-02 1.04e-02 3.04e+00
54 2.90e+01 5.41e+00 4.51e+02 2.10e+01 0.00e+00 1.27e-02 1.00e-02 3.04e+00
55 2.89e+01 5.41e+00 4.49e+02 2.00e+01 0.00e+00 1.23e-02 9.76e-03 3.04e+00
56 2.87e+01 5.40e+00 4.48e+02 1.90e+01 0.00e+00 1.20e-02 9.54e-03 3.04e+00
57 2.86e+01 5.40e+00 4.46e+02 1.80e+01 0.00e+00 1.16e-02 9.27e-03 3.04e+00
58 2.85e+01 5.40e+00 4.44e+02 1.71e+01 0.00e+00 1.13e-02 8.94e-03 3.04e+00
59 2.84e+01 5.40e+00 4.43e+02 1.63e+01 0.00e+00 1.10e-02 8.66e-03 3.04e+00
60 2.82e+01 5.40e+00 4.41e+02 1.56e+01 0.00e+00 1.06e-02 8.40e-03 3.04e+00
61 2.82e+01 5.40e+00 4.40e+02 1.49e+01 0.00e+00 1.03e-02 8.20e-03 3.04e+00
62 2.81e+01 5.40e+00 4.39e+02 1.42e+01 0.00e+00 9.98e-03 7.96e-03 3.04e+00
63 2.80e+01 5.40e+00 4.38e+02 1.36e+01 0.00e+00 9.70e-03 7.75e-03 3.04e+00
64 2.79e+01 5.40e+00 4.37e+02 1.30e+01 0.00e+00 9.40e-03 7.56e-03 3.04e+00
65 2.78e+01 5.40e+00 4.36e+02 1.25e+01 0.00e+00 9.12e-03 7.41e-03 3.04e+00
66 2.77e+01 5.39e+00 4.35e+02 1.19e+01 0.00e+00 8.87e-03 7.23e-03 3.04e+00
67 2.76e+01 5.39e+00 4.34e+02 1.14e+01 0.00e+00 8.58e-03 7.08e-03 3.04e+00
68 2.76e+01 5.39e+00 4.32e+02 1.09e+01 0.00e+00 8.32e-03 6.91e-03 3.04e+00
69 2.75e+01 5.39e+00 4.31e+02 1.05e+01 0.00e+00 8.12e-03 6.66e-03 3.04e+00
70 2.74e+01 5.39e+00 4.30e+02 1.01e+01 0.00e+00 7.88e-03 6.51e-03 3.04e+00
71 2.73e+01 5.39e+00 4.29e+02 9.68e+00 0.00e+00 7.66e-03 6.33e-03 3.04e+00
72 2.73e+01 5.39e+00 4.28e+02 9.33e+00 0.00e+00 7.45e-03 6.19e-03 3.04e+00
73 2.72e+01 5.39e+00 4.27e+02 8.99e+00 0.00e+00 7.25e-03 6.04e-03 3.04e+00
74 2.72e+01 5.39e+00 4.27e+02 8.69e+00 0.00e+00 7.05e-03 5.89e-03 3.04e+00
75 2.71e+01 5.39e+00 4.26e+02 8.39e+00 0.00e+00 6.88e-03 5.72e-03 3.04e+00
76 2.70e+01 5.39e+00 4.25e+02 8.11e+00 0.00e+00 6.71e-03 5.61e-03 3.04e+00
77 2.70e+01 5.39e+00 4.24e+02 7.83e+00 0.00e+00 6.52e-03 5.51e-03 3.04e+00
78 2.69e+01 5.39e+00 4.23e+02 7.56e+00 0.00e+00 6.36e-03 5.39e-03 3.04e+00
79 2.69e+01 5.39e+00 4.22e+02 7.32e+00 0.00e+00 6.53e-03 5.25e-03 2.77e+00
80 2.68e+01 5.39e+00 4.22e+02 7.11e+00 0.00e+00 6.41e-03 5.13e-03 2.77e+00
81 2.68e+01 5.39e+00 4.21e+02 6.91e+00 0.00e+00 6.28e-03 5.00e-03 2.77e+00
82 2.68e+01 5.39e+00 4.21e+02 6.72e+00 0.00e+00 6.13e-03 4.88e-03 2.77e+00
83 2.67e+01 5.39e+00 4.21e+02 6.54e+00 0.00e+00 6.02e-03 4.73e-03 2.77e+00
84 2.67e+01 5.39e+00 4.20e+02 6.35e+00 0.00e+00 5.90e-03 4.65e-03 2.77e+00
85 2.67e+01 5.39e+00 4.20e+02 6.17e+00 0.00e+00 5.77e-03 4.55e-03 2.77e+00
86 2.66e+01 5.39e+00 4.19e+02 5.99e+00 0.00e+00 5.66e-03 4.47e-03 2.77e+00
87 2.66e+01 5.39e+00 4.19e+02 5.82e+00 0.00e+00 5.54e-03 4.41e-03 2.77e+00
88 2.66e+01 5.39e+00 4.18e+02 5.65e+00 0.00e+00 5.45e-03 4.32e-03 2.77e+00
89 2.65e+01 5.39e+00 4.17e+02 5.49e+00 0.00e+00 5.33e-03 4.25e-03 2.77e+00
90 2.65e+01 5.39e+00 4.17e+02 5.33e+00 0.00e+00 5.22e-03 4.19e-03 2.77e+00
91 2.65e+01 5.39e+00 4.16e+02 5.18e+00 0.00e+00 5.12e-03 4.13e-03 2.77e+00
92 2.64e+01 5.39e+00 4.16e+02 5.04e+00 0.00e+00 5.01e-03 3.94e-03 2.77e+00
93 2.64e+01 5.39e+00 4.15e+02 4.92e+00 0.00e+00 4.90e-03 3.84e-03 2.77e+00
------------------------------------------------------------------------------------
ConvBPDN solve time: 42.27s
Reconstruct image from sparse representation.
shr = b.reconstruct().squeeze()
imgr = sl + shr
print("Reconstruction PSNR: %.2fdB\n" % sm.psnr(img, imgr))
Reconstruction PSNR: 37.30dB
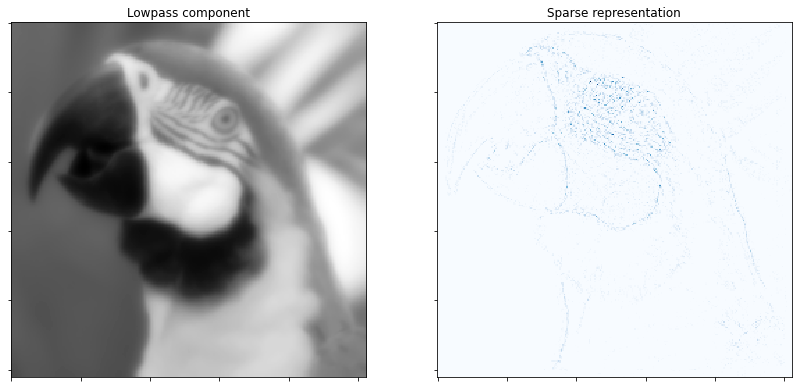
Display low pass component and sum of absolute values of coefficient maps of highpass component.
fig = plot.figure(figsize=(14, 7))
plot.subplot(1, 2, 1)
plot.imview(sl, title='Lowpass component', fig=fig)
plot.subplot(1, 2, 2)
plot.imview(np.sum(abs(X), axis=b.cri.axisM).squeeze(), cmap=plot.cm.Blues,
title='Sparse representation', fig=fig)
fig.show()

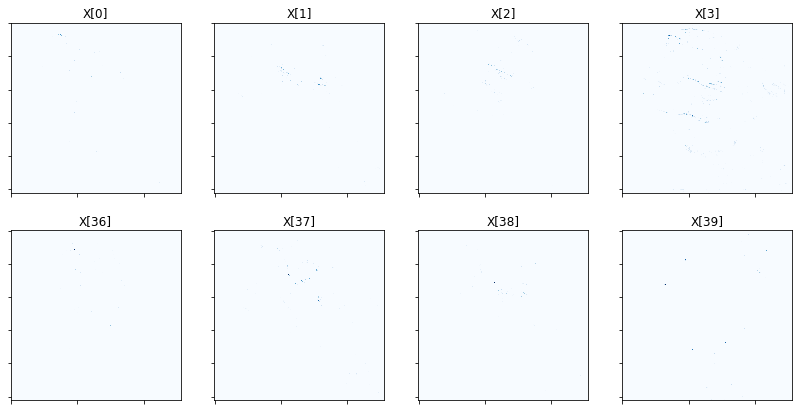
Show activation of grouped elements column-wise for first four groups. As mu is lowered, the vertical pairs should look more and more similar. You will likely need to zoom in to see the activations clearly.
fig = plot.figure(figsize=(14, 7))
for i in range(4):
plot.subplot(2, 4, i + 1)
plot.imview(abs(X[:, :, :, :, i]).squeeze(), cmap=plot.cm.Blues,
title=f'X[{i}]', fig=fig)
plot.subplot(2, 4, i + 5)
plot.imview(abs(X[:, :, :, :, i + 36]).squeeze(), cmap=plot.cm.Blues,
title=f'X[{i+36}]', fig=fig)
fig.show()

Display original and reconstructed images.
fig = plot.figure(figsize=(14, 7))
plot.subplot(1, 2, 1)
plot.imview(img, title='Original', fig=fig)
plot.subplot(1, 2, 2)
plot.imview(imgr, title='Reconstructed', fig=fig)
fig.show()

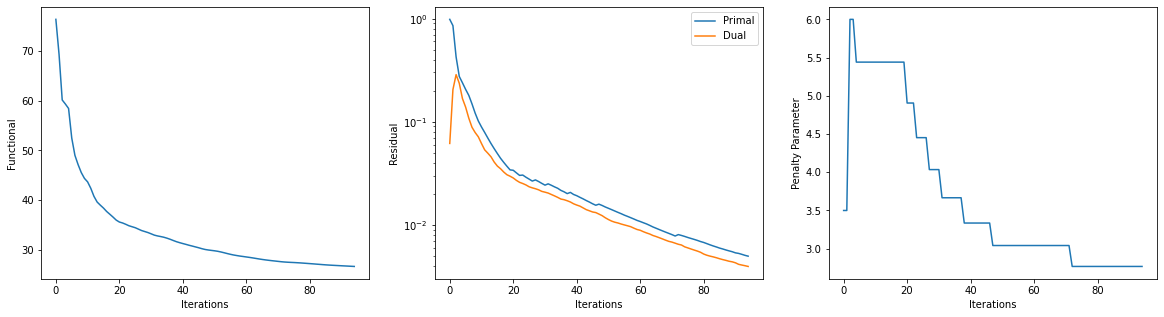
Get iterations statistics from solver object and plot functional value, ADMM primary and dual residuals, and automatically adjusted ADMM penalty parameter against the iteration number.
its = b.getitstat()
fig = plot.figure(figsize=(20, 5))
plot.subplot(1, 3, 1)
plot.plot(its.ObjFun, xlbl='Iterations', ylbl='Functional', fig=fig)
plot.subplot(1, 3, 2)
plot.plot(np.vstack((its.PrimalRsdl, its.DualRsdl)).T,
ptyp='semilogy', xlbl='Iterations', ylbl='Residual',
lgnd=['Primal', 'Dual'], fig=fig)
plot.subplot(1, 3, 3)
plot.plot(its.Rho, xlbl='Iterations', ylbl='Penalty Parameter', fig=fig)
fig.show()